In this article, MedChem Express reviews the evolution and pipeline of personalized cancer vaccines, highlighting neoantigen sources, screening technologies, and key translational challenges.
Personalized neoantigen vaccines target unique mutations in individual tumors, activating potent and specific T‑cell responses while minimizing the risk of autoimmunity. Recent breakthroughs in next-generation sequencing (NGS), immunopeptidomics, AI-powered prediction algorithms, and diverse vaccine platforms (e.g., mRNA/LNP) have enabled the precise identification of high-quality neoantigens from vast mutational landscapes.
These advances, supported by research tools and compound libraries from MedChem Express, facilitate the rapid translation of candidate neoantigens into clinical-grade vaccines. The positive Phase III results of an mRNA-based neoantigen vaccine in adjuvant melanoma (mRNA-4157) further validate this approach as a transformative clinical modality[1].
Evolution and Pipeline of Personalized Cancer Vaccines
The development of cancer vaccines has evolved from broadly targeting tumor-associated antigens (TAAs) toward precise, individualized targeting of tumor-specific antigens (TSAs), particularly neoantigens.
Evolution of Cancer Vaccines: From Shared Antigens to Personalized Neoantigen Vaccines
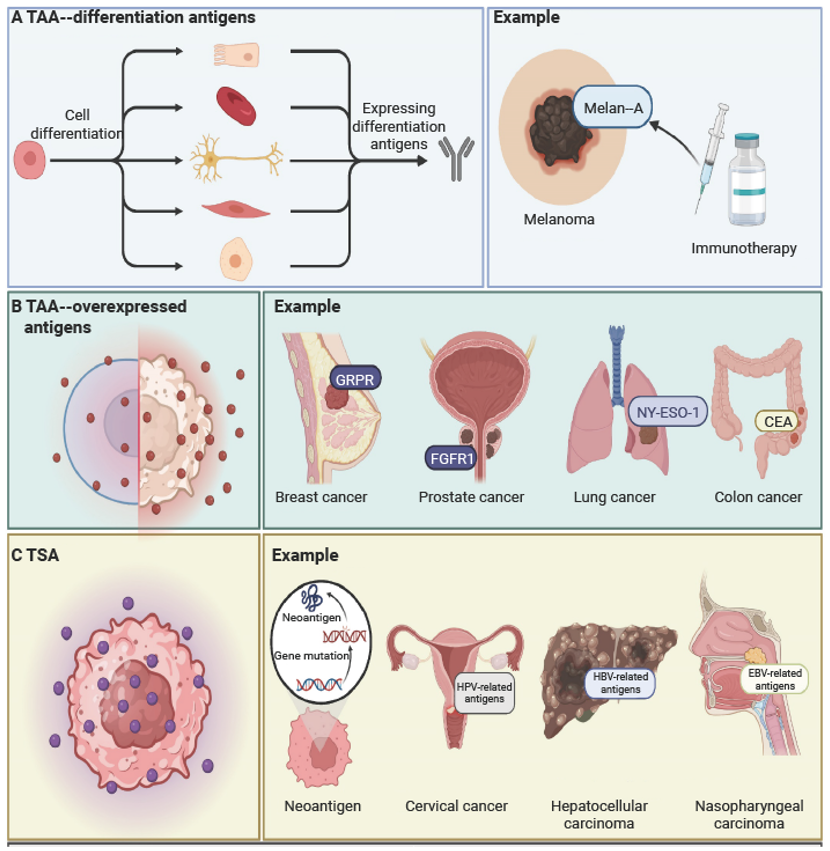
- Tumor-associated antigens (TAAs): TAAs are “self-antigens” abnormally expressed in tumors, encompassing overexpressed proteins, cancer-germline antigens, and differentiation antigens. They are often shared across patients and cancer types, facilitating large-scale vaccine production[2]. However, their development is limited by central thymic tolerance, which depletes high-affinity T cells against self-peptides, and carries the risk of on-target/off-tumor toxicity against normal tissues. Consequently, early TAA-focused strategies using protein, DNA, viral vector, or whole-cell platforms showed limited clinical efficacy, hampered by pre-existing immune tolerance, suboptimal adjuvants, and the immunosuppressive tumor microenvironment (TME)[3].
- Tumor-specific antigens (TSAs): TSAs, including viral oncoproteins and neoantigens, provide high tumor specificity and immunogenicity. Examples include the E6/E7 proteins of human papillomavirus (HPV). Preventive vaccines based on such antigens, like HPV vaccines, have achieved significant success in reducing the incidence of cervical and related cancers[4]. However, their application is inherently limited to virus-associated malignancies and cannot address the majority of solid tumors.
- Neoantigens: Neoantigens, a major subclass of TSAs, are unique peptides arising from gene mutations in tumor cells. They are recognized as “non-self” by the immune system, effectively bypassing immune tolerance and eliciting robust anti-tumor T-cell responses[5-6]. Their highly individualized nature poses challenges for vaccine development in terms of complexity and cost. Personalized neoantigen vaccines aim to leverage this specificity to overcome tumor heterogeneity while minimizing off-target toxicities, driving broad and durable anti-tumor immunity. Early-phase clinical trials underscore this potential: two 2025 studies demonstrated efficacy in renal cell carcinoma (n=9)[7] and pancreatic ductal adenocarcinoma (n=8)[8] using vaccines targeting patient-specific mutations.
Development Process of Personalized Cancer Vaccines
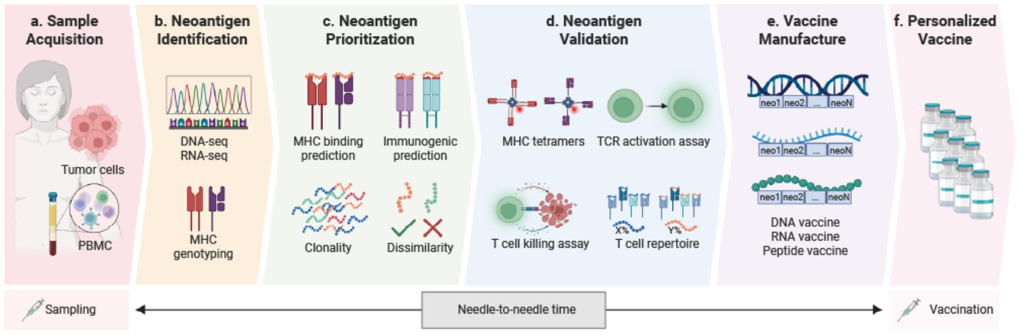
The development of personalized cancer vaccines is a complex, multi-step process that integrates cutting-edge genomics, bioinformatics, and immunology technologies[9]. The core workflow includes:
- Tumor Sample Analysis & Neoantigen Identification
Biopsy samples are obtained from the patient’s tumor, with matched normal tissue (e.g., PBMCs) serving as a germline control. NGS of DNA and RNA is performed to identify tumor-specific genetic mutations. Immunopeptidomics can complement this by detecting mutation-independent, cryptic neoantigens arising from aberrant transcription or translation. The sequencing data are also used for Major Histocompatibility Complex (MHC) genotyping of the patient. - Neoantigen Screening & Prioritization
Bioinformatics algorithms prioritize candidate neoantigens based on key filters: predicted MHC binding affinity, immunogenic potential for T-cell activation, clonality (favoring clonal neoantigens present in most tumor cells to prevent immune escape), and dissimilarity from self-peptides to ensure safety. - In Vitro Neoantigen Validation
Top-priority candidates undergo functional validation assays, bridging prediction and clinical application. Key methods include synthesizing MHC tetramers loaded with neoantigenic peptides to identify and quantify specific T cells via flow cytometry, as well as conducting T-cell activation assays (e.g., ELISpot) and cytotoxicity assays to confirm their ability to robustly activate T cells and kill antigen-presenting target cells. - Vaccine Design & Synthesis
Based on the final neoantigen epitopes, a personalized vaccine is custom-synthesized, most commonly as a multi-peptide or mRNA vaccine. The formulation often includes adjuvants or uses delivery platforms such as nanoparticles to enhance immunogenicity and delivery efficiency. Minimizing the total “needle-to-needle” time—from biopsy to finished vaccine—is crucial for timely therapeutic intervention. - Clinical Application & Evaluation
The vaccine is administered to the patient, often in combination with other therapies such as immune checkpoint inhibitors (ICIs) to improve outcomes. Efficacy is evaluated by monitoring the expansion of neoantigen-specific T cells and clinical response.
The shift from broadly shared TAAs to highly specific TSAs highlights the importance of neoantigen discovery. In the following section, we explore the diverse sources of neoantigens and the innovative screening approaches that translate these discoveries into actionable therapeutic candidates.
Neoantigen Sources and Screening Methods
Sources of Neoantigens
Neoantigens primarily originate from tumor-specific genomic alterations absent in normal cells, making them highly tumor-specific and recognizable as “non-self” by the immune system[10]. Traditionally, neoantigens have been categorized based on canonical genomic and transcriptomic variations. However, recent advances in immunopeptidomics and deep sequencing have unveiled a broader repertoire of “hidden” or cryptic neoantigens derived from genomic regions previously considered non-coding. These two paradigms, canonical and non-canonical, together define the modern understanding of neoantigen sources.
1. Canonically Neoantigens
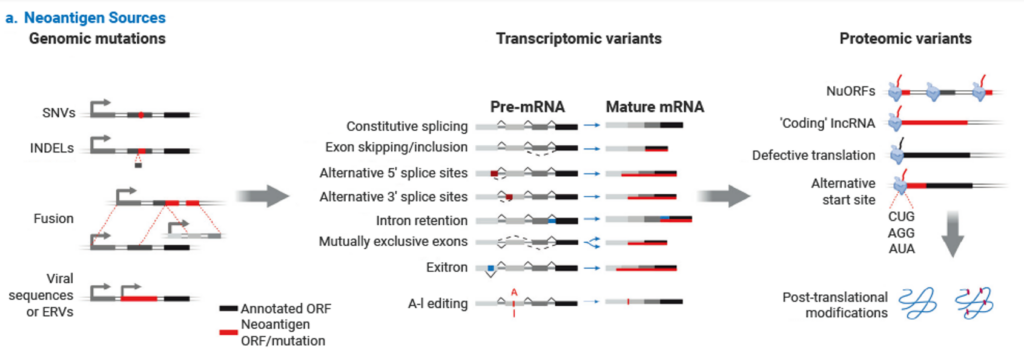
- Genomic mutations: Somatic alterations, including single-nucleotide variants (SNVs), insertions and deletions (INDELs), and gene fusions, are primary drivers of neoantigen production. These mutations alter the amino acid sequence, generating novel peptides presented by MHC molecules.
- Aberrant transcriptomic variants: RNA-level processing errors, including alternative splicing, polyadenylation, and RNA editing (e.g., A-to-I editing), can create novel transcript isoforms and consequently new peptide sequences.
- Post-translational modifications (PTMs): Altered or tumor-specific PTMs (e.g., phosphorylation, citrullination) can generate novel epitopes by modifying the structure or charge of presented peptides, even without changes to the underlying DNA sequence.
- Viral open reading frames (ORFs): In virus-associated cancers (e.g., HPV, EBV), proteins encoded by viral ORFs constitute foreign, highly immunogenic antigens.
2. “Hidden” Neoantigens
This emerging category arises not from simple sequence mutations, but from the aberrant translation of genomic regions traditionally ignored by standard annotation. Zackery et al. systematically discovered a vast repertoire of “cryptic antigens” derived from non-coding regions in pancreatic cancer and demonstrated their ability to induce tumor-killing T-cell responses[12].
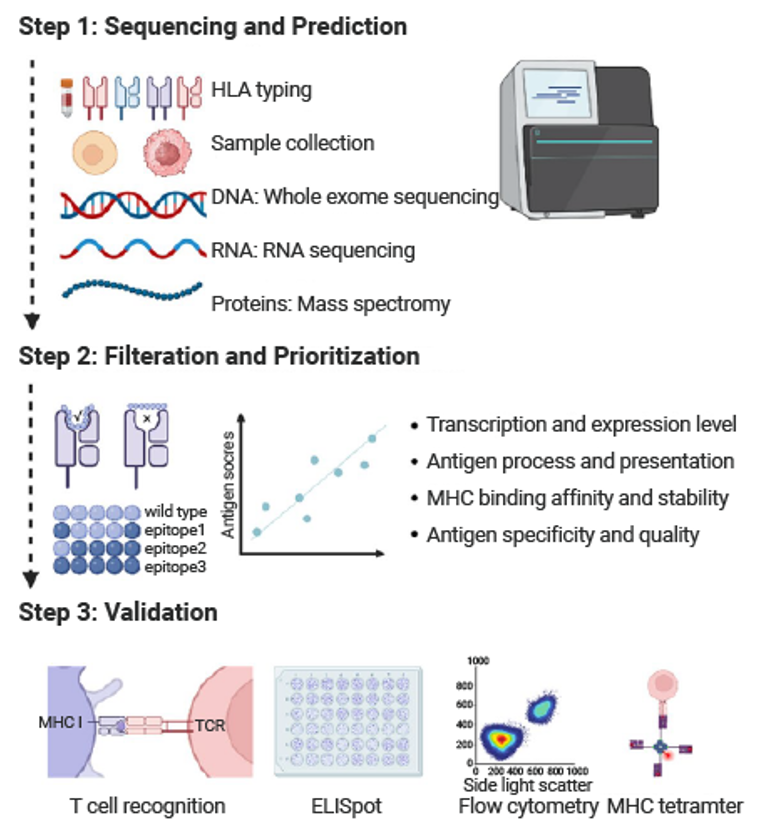
- Prediction: This initial stage involves identifying tumor-specific mutations through sequencing of tumor DNA, RNA, and protein sequencing, correlated with the patient’s HLA typing. Computational tools are employed to detect these mutations and predict which are likely to generate peptides recognizable by the immune system.
- Filtration and Prioritization: After predicting numerous potential neoantigens, a rigorous filtration process is undertaken to rank and prioritize these candidates. Features such as expression levels, likelihood of processing and presentation on MHC molecules, MHC binding affinity, and antigen specificity are assessed. This step aims to select the most promising neoantigens capable of eliciting a strong immune response.
- Validation: The final step involves experimental methods to confirm the selected neoantigens’ ability to elicit specific T-cell responses. Techniques such as Enzyme-Linked Immunospot (ELISpot) assays can detect activation and cytokine secretion of neoantigen-specific T cells, thereby validating their immunogenicity.
Artificial Intelligence (AI) in Neoantigen Screening
Machine learning and AI play a crucial role in neoantigen screening by significantly enhancing the efficiency, accuracy, and breadth of the process, thereby accelerating the development of personalized cancer immunotherapy[13]. AI technologies can process and analyze vast amounts of biological data, identify potential neoantigens that are difficult to detect using traditional methods, and optimize predictions of their immunogenicity.
For instance, Cai et al. developed MaNeo, an immunopeptidomics-guided screening pipeline using Random Forest (RF) and XGBoost classifiers[14]. Functional validation demonstrating that the prioritized neo-peptides effectively induce T cell proliferation and specific cytotoxicity against cancer cells without harming healthy cells. This approach enables more comprehensive and accurate discovery of both canonical and non-canonical neoantigens, thereby broadening the target pool for personalized cancer immunotherapy. The next frontier involves deep learning architectures capable of modeling long-range dependencies in peptide-MHC interactions, as well as algorithms that integrate single-cell TCR sequencing data to predict neoantigen immunogenicity based on a patient’s pre-existing T-cell repertoire.
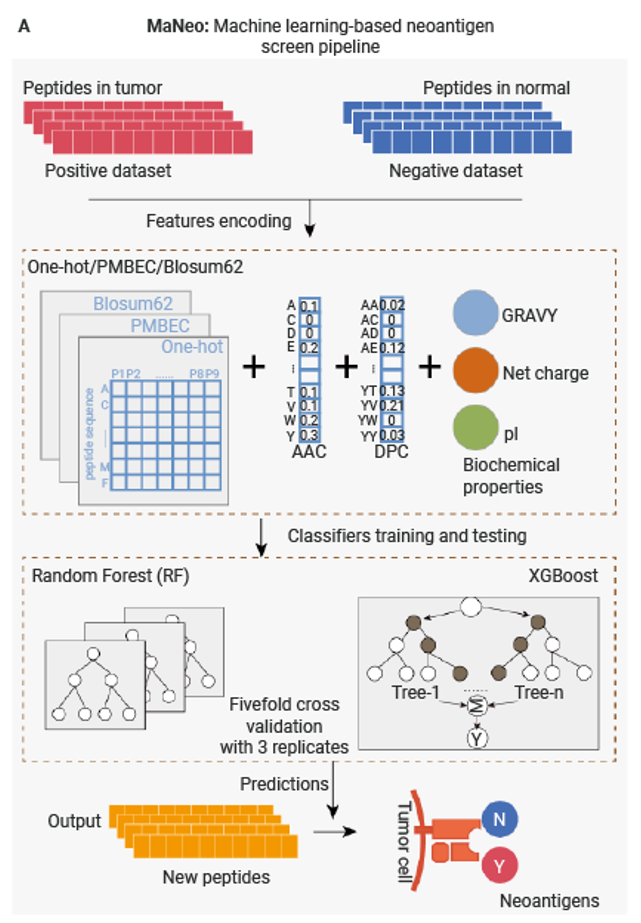
Collectively, these approaches, predicting neoantigen immunogenicity, filtering and prioritizing candidates, validating T-cell responses, and leveraging AI to optimize predictions, form a robust framework for identifying clinically relevant neoantigens.
Vaccine Platform Selection and Translational Challenges
Platform Selection of Personalized Cancer Vaccines
Following the successful identification of highly immunogenic neoantigens, the focus shifts to a critical translational bottleneck: the efficient delivery of these antigens and the induction of a robust and durable immune response. The choice of vaccine platform is therefore paramount, as it directly influences delivery efficiency, immunogenicity, safety, and ultimately clinical success.
Table 1 summarizes the core advantages, limitations, and mechanisms of the leading vaccine platform technologies. To bridge the gap between preclinical promise and clinical reality, we highlight key innovative studies alongside emerging clinical trial data for each approach. Given its transformative potential, mRNA-based neoantigen vaccination is discussed in a dedicated subsection.
| Vaccine platform | Advantages | Limitations | Examples |
| Peptide | Easy of production; Minimal toxicity; High specificity and safety | Limited immunogenicity; Short half-life; HLA restriction | OSE2101, IO101 |
| DNA | Cost-effectiveness; Stable; Durable immunity | Risk of gene integration; Limited immunogenicity; Low transfection efficacy | GX-188E, VGX-3100 |
| mRNA | Flexibly design; Rapid production; Potent immune activation | Instability; Inefficient delivery; Susceptible to degradation | BNT111, mRNA-4157 (V940) |
| Replication-defective viruses | High immunogenicity; High delivery efficiency | Pre-existing immunity; Risk of unintended viral spread | TG4010, TroVax |
| Virus-like particles | High immunogenicity; No risk of infection; Stable and Scalable production | Limited T-cell activation; Formulation issues | Gardasil 9, ES2B-C001, CMP-001 |
| Oncolytic viruses | Direct tumor lysis; Potent immune activation; Strong tumor targeting | Safety risks; Immune clearance; Complex production | T-VEC, RP-1, JX-594 |
| Tumor cells | Broad antigen coverage; Reduced off-target effects | Risk of tumorigenicity; Complex production | GVAX, M-Vax, Canvaxin |
| Dendritic cells | Effective antigen presentation; Strong T-cell activation | High cost; Complex production | Sipuleucel-T, Ilixadencel |
Core Bottlenecks and Solutions for Therapeutic Neoantigen Vaccines
Following the discussion on platform technologies, it is important to recognize that translating neoantigen vaccines into effective therapies faces several fundamental biological and technical bottlenecks.
The development of therapeutic neoantigen vaccines encounters three major challenges[11]:
- Limited prediction accuracy: Precisely identifying the few immunogenic neoantigens among a vast number of tumor mutations that are actually recognized by the immune system is extremely challenging. Only a tiny fraction (~1%) of algorithm-predicted candidates truly activate T cells, making this the critical rate-limiting step for personalized therapy.
- Tumor immune evasion: Tumors can actively lose neoantigens through gene loss or epigenetic silencing, or downregulate MHC molecules to impair antigen presentation. In addition, the immunosuppressive tumor microenvironment further compromises T-cell function.
- Insufficient neoantigen-specific T-cell response: The pre-existing repertoire of neoantigen-reactive T cells in patients is often limited. Moreover, T cells obtained for adoptive cell therapies may also exhibit poor persistence and terminal differentiation, undermining therapeutic efficacy.
To address these challenges, integrated strategies are being developed. Efforts focused on improving neoantigen prediction by combining genomics with experimental immunopeptidomics, countering immune evasion through combination therapies with checkpoint inhibitors or “off-the-shelf” public neoantigen vaccines, and enhancing T-cell responses via optimized expansion protocols or engineered cell therapies such as TCR-T cells[15].
Summary
Cancer vaccine development has shifted from broadly shared tumor-associated antigens to patient-specific neoantigens, which arise from diverse sources such as genomic mutations, transcriptomic alterations, post-translational modifications, and cryptic non-coding regions. These neoantigens provide highly individualized targets for potent anti-tumor immune responses, though their effectiveness relies on efficient delivery and robust immune activation. Various vaccine platforms, including peptide, DNA, mRNA, viral, and cell-based systems, facilitate this delivery but still face challenges related to immunogenicity, safety, and tumor immune evasion.
Looking ahead, parallel advances in neoantigen identification, vaccine platform engineering, and immune combination strategies are expected to make personalized neoantigen vaccines a pivotal component of the immunotherapy landscape. This integrated approach promises greater precision and durability in anti-tumor responses, offering new hope for patients.
Recommended Products and Compound Libraries
| Category | Product Name | Cat. No. | Description |
| Tools for T Cell Activation / Functional Assays | |||
| Recombinant Protein | IL-2 Protein, Human (His) | HY-P7037B | Produced by activated CD4+ helper T cells; important for immune response and tolerance. |
| IL-7 Protein, Human (HEK293, His) | HY-P70755 | Hematopoietic cytokine essential for development, expansion, and survival of T and B cells. | |
| Antibody | Anti-CD3 Antibody (OKT-3) | HY-P990864 | Mouse IgG2a κ antibody targeting human CD3; inhibits T cells; for immunology research. |
| Anti-CD28 Antibody (9.3) | HY-P990865 | Anti-human CD28 IgG2a monoclonal antibody; activates T cells; for immunology research. | |
| MCE Kits | Human CD3/CD28 T Cell Activation Magnetic Beads | HY-K0353 | Provides CD3/CD28 co-stimulation without feeder cells or antigens; enables simple, rapid T cell activation. |
| Mouse CD3/CD28 T Cell Activation Magnetic Beads | HY-K0354 | Provides CD3/CD28 co-stimulation without feeder cells or antigens; enables simple, rapid T cell activation. | |
| Compound Libraries for Cancer Research | |||
| Compound Library | Anti-Cancer Compound Library | HY-L025 | Compound library containing 9900+ small molecules with anti-cancer activity. |
| Anti-Cancer Natural Product Library | HY-L107 | Compound library containing 1800+ natural products with identified orpotential anti-cancer activity. | |
| FDA-Approved Anticancer Drug Library | HY-L122 | A unique collection of 1400+ approved drugs with anti-cancer activity. | |
| Small Molecule Immuno-Oncology Compound Library | HY-L031 | Compound library containing 690+ targeted small molecules in immuno-oncology. | |
References
[1] Weber JS, et al. Lancet. 2024 Feb 17;403(10427):632-644.
[2] Lin P, et al. Theranostics. 2025 Jan 1;15(1):300-323.
[3] Peng K, et al. Cell Mol Immunol. 2025 Aug;22(8):840-868.
[4] BioNTech SE. ClinicalTrials.gov, 2025-12-10.
[5] Baharom F, et al. Nat Rev Immunol. 2025 Mar;25(3):195-211.
[6] Zhou Y, et al. J Hematol Oncol. 2025 Feb 17;18(1):18.
[7] Braun DA, et al. Nature. 2025 Mar;639(8054):474-482.
[8] Sethna Z, et al. Nature. 2025 Mar;639(8056):1042-1051.
[9] Chi WY, et al. J Biomed Sci. 2024 Oct 9;31(1):94.
[10] Han Y, et al. Nature. 2026 Jan 7.
[11] Xie N, et al. Signal Transduct Target Ther. 2023 Jan 6;8(1):9.
[12] Ely ZA, et al. Science. 2025 May 8;388(6747):eadk3487.
[13] Zhong Q, et al. Brief Bioinform. 2025 Jul 2;26(4):bbaf254.
[14] Cai Y, et al. Sci Adv. 2025 May 23;11(21):eadv6445.
[15] Katsikis PD, et al. Nat Rev Immunol. 2024 Mar;24(3):213-227.
Inhibitors | Agonists | Recombinant Proteins | Dyes | Assay Kits | Compound Libraries | Drug Screening | PCR | Cell Culture Reagents | Custom Synthesis, Screening, ADC, PROTAC Solutions