Wide range of CRISPR knockout and activation products
Santa Cruz Biotechnology offers target-specific CRISPR/Cas9 knockout plasmids, CRISPR double nickase plasmids, CRISPR/ dCas9 activation plasmids and CRISPR lenti activation systems for over 18,910 human and 18,340 mouse protein encoding genes.
The CRISPR/Cas system is an adaptive immune defense mechanism used by Archea and bacteria for the degradation of foreign genetic material. In these organisms, the foreign genetic material from a bacteriophage is acquired and integrated into the CRISPR loci (1,2). This new material, also known as a spacer, creates a sequence-specific fragment used for future resistance against a bacteriophage infection. These sequence-specific fragments are translated into short CRISPR RNAs (crRNAs) and function as a guide to direct cleavage of complementary invading DNA via the nuclease activity of CRISPR-associated (Cas) protein also encoded by the CRISPR loci (1,2). Cas9 nuclease of the type II CRISPR system has an RNA binding domain, an alpha helix recognition lobe (REC), a nuclease lobe that include the RuvC and HNH for DNA cleavage, and a protospacer adjacent motif (PAM) interacting site (1,2). crRNA forms a complex with the Cas9 nuclease by binding to the bridge helix within the REC lobe, and forms multiple salt bridges with the backbone of the crRNA (1,2,3).
Once the crRNA binds to the Cas9 the conformation of the Cas9 nuclease changes and creates a channel that allows for DNA binding (1,2,3). The Cas9/crRNA complex scans the DNA for a PAM (5′-NGG) site (4,5,6). Recognition of a PAM site leads to unwinding of the DNA, and allows the crRNA to check for complementary DNA adjacent to the PAM site. When Cas9 binds to a PAM site adjacent to a DNA sequence that is complementary to the crRNA, the bridge helix within the REC lobe creates an RNA-DNA heteroduplex with the target DNA (3,4,7). The PAM site recognition is involved in activating the nucleolytic HNH and RuvC domains which create a double-stranded break (DSB) in the target DNA, leading to DNA degradation (1,2,5,8). If the crRNA is not complementary, then Cas9 releases and searches for another PAM site (7). Targeted genome strand breaks in the DNA can be repaired via the nonhomologous end-joining (NHEJ) repair pathway, which introduces insertion or deletions creating errors, or through the homologous directed repair (HDR) pathway, that can be used to recombine selected markers at specific sites in the genome (2,9,10). This CRISPR/Cas9 mechanism can be repurposed for genomic engineering of various systems, including mammalian cells.
Genome editing via introduction of DSBs can be performed with meganucleases, zinc finger nucleases (ZF), or transactivator-like effectors (TALEs), which recognize DNA sequences, however, each has their limitations. When using meganuclesases it is difficult to clearly show site-specific recognition between nuclease and DNA (2).The other options, ZFs and TALEs, have proven difficult to design and recognize up to 3 nt of DNA (2). Single guide RNAs (sgRNAs) that act like crRNAs are easily designed and can be expressed along with Cas9 nuclease in the same vector to target specific DNA sites for genome editing. The CRISPR/Cas 9 system also has higher sensitivity and is more efficient when used for screening than small hairpin RNAs.
CRISPR/Cas9-directed DSB
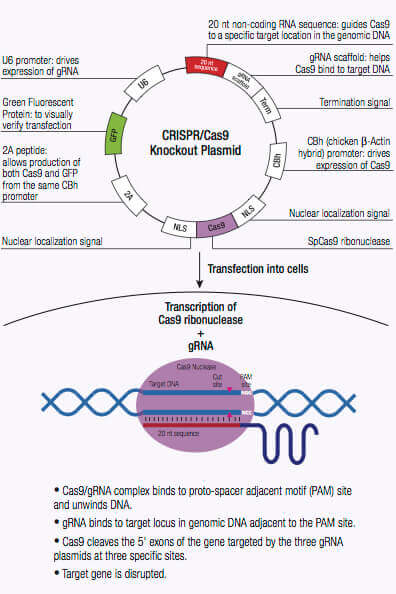
CRISPR/Cas9 plasmids
- 20 µg, up to 20 transfections
- CRISPR/Cas9 knockout plasmids consist of a pool of three plasmids each encoding the Cas9 nuclease and a target-specific 20 nt guide RNA (gRNA) designed for maximum knockout efficiency
- gRNA sequences are derived from the GeCKO (v2) library and direct the Cas9 protein to induce a site-specific double strand break (DSB) in the genomic DNA (11)
- CRISPR/Cas9 knockout plasmids available for human and mouse genes are indicated by the (h) or (m) designation in the product name. Example: p53 CRISPR/Cas9 KO Plasmids (h) for human or p53 CRISPR/Cas9 KO Plasmids (m) for mouse
- Provided as transfection-ready, purified plasmid DNA
Support products for CRISPR/Cas9 plasmids
- Suitable control antibodies are available
- Ultracruz Transfection Reagent (sc-395739)
- Plasmid Transfection Medium (sc-108062)
- Control CRISPR/Cas9 Plasmid (sc-418922)
Tumor suppressor/apoptosis
Synthesis and degradation gene silencers
Growth factors and hormone gene silencers
Neurobiology gene silencers
HDR plasmids
HDR plasmids provide a specific DNA repair template for a DSB, and are only used when co-transfected with CRISPR/Cas9 KO plasmids. When co-transfected with the CRISPR/ Cas9 KO plasmid, the HDR plasmid incorporates a puromycin resistance gene for selection of cells where Cas9-induced DNA cleavage has occurred.
HDR plasmid product details
- 20 µg, up to 20 transfections
- Target specific HDR Plasmids are recommended for co-transfection with the CRISPR/Cas9 KO plasmid for the same gene target and species
- HDR plasmid consists of a pool of 3 plasmids, each containing a homology-directed DNA repair (HDR) template corresponding to the cut sites generated by the corresponding CRISPR/Cas9 KO plasmids
- Each HDR template contains two 800 bp homology arms designed to specifically bind to the genomic DNA surrounding the corresponding Cas9-induced double strand DNA break site
- Each HDR plasmid inserts a puromycin resistance gene to enable selection of stable knockout (KO) cells
- Each puromycin resistance gene is flanked by two LoxP sites to allow for further processing by the Cre vector
- Each HDR plasmid also contains RFP to visually confirm transfection
- Provided as transfection-ready, purified plasmid DNA
Support products for HDR plasmids
- Ultracruz Transfection Reagent (sc-395739)
- Plasmid Transfection Medium (sc-108062)
- Puromycin dihydrochloride (sc-108071)
Transcription regulator gene silencers
Signaling intermediate gene silencers
Transport and trafficking gene silencers
Membrane receptor gene silencers
Cre Vector
The Cre Vector expresses Cre recombinase, a bacteriophage p1 enzyme that catalyzes site-specific DNA recombination between two LoxP sites. When the CRISPR/Cas9 knockout plasmid is co- transfected with the HDR plasmid, cells containing the edited DNA can be isolated using the selection marker inserted during homology-directed repair. Following selection, cells can be transfected with the Cre Vector to excise the genetic material inserted during homology-directed repair, such as the puromycin resistance gene.
Cre Vector product details
- 20 µg, up to 20 transfections
- Recommended for DNA repair of selected cells successfully edited by CRISPR/Cas9 KO plasmid and HDR plasmid
- Cre Vector contains a CMV promoter to drive expression of Cre recombinase
- Provided as a transfection-ready, purified plasmid DNA
- Cre Vector (sc-418923)
Support products for Cre Vector
Looking for other CRISPR/Cas9 knockout plasmids?
These are the CRISPR/Cas9 knockout plasmids categories Santa Cruz Biotechnology offer:
- Tumor suppressors/apoptosis
- Cell cycle proteins
- Transcription regulators
- Homeodomain proteins
- Steroid receptors
- Kinases and phosphatases
- Growth factors and hormones
- Membrane receptors
- Signaling intermediates
- Transport and trafficking
- Synthesis and degradation
- Neurobiology
- Channel proteins
- Gdp/gtp binding proteins
- Lymphocyte signaling
- Cell adhesion proteins
- Structural proteins
- Additional gene products
References
- Van der Oost J., et al. 2014. Unraveling the Structural and Mechanistic Basis of CRISPR-Cas Systems. Nat. Rev. Microbiol. (7):479-92. PMID 24909109.
- Hsu, P., et al. 2014. Development and Applications of CRISPR-Cas9 for Genome Editing. Cell. 157(6):1262-78. PMID 24906146.
- Nishimasu, H., et al. 2014. Crystal Structure of Cas9 in Complex with Guide RNA and Target DNA. Cell. 156(5):935-49. PMID 24529477.
- Deltcheva, E., et al. 2011. CRISPR RNA Maturation by Trans-Encoded Small RNA and Host Factor RNASE III. Nature. 471: 602-607. PMID 21455174.
- Jinek, M., et al. 2012. A Programmable Dual-RNA Guided DNA Endonuclease in Adaptive Immunity. Science. 337(6096):816-21. PMID 22745249.
- Deveau, H., et al. 2010. CRISPR/Cas System and its Role in Phage-Bacteria Interaction. Annu. Rev. Microbiol. 64: 475-493. PMID 20528693.
- Sternberger, SH., et al. 2014. DNA Interrogation by the CRISPR RNA-guided Endonuclease Cas9. Nature. 507(7490):62-7. PMID 24476820.
- Gasuinas, G., et al. 2012. Cas9-crRNA Ribonucleoprotein Complex Mediates Specific DNA Cleavage for Adaptive Immunity in Bacteria. Proc. Natl. Acad. Sci. 109(39):E2579-86. PMID 22949671.
- Mali, P., et al. 2013. RNA-Guided Human Genome Engineering via Cas9. Science. 339 (6121): 823-6. PMID 23287722.
- Ran, FA., et al. 2013. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. (11): 2281-308. doi: 10.1038/nprot.2013.143. PMID 24157548.
- Shalem O., et al. 2014. Genome-scale CRISPR-Cas9 Knockout Screening in Human Cells. 343(6166):84-7. PMID 24336571.

Monoclonal Antibodies | Secondary Antibodies | ChemCruz Biochemicals | Labware | CRISPR | Gene Editing







